Our Science
We Build Nanobots
Using the magic of DNA hybridization we design and build self-assembling
DNA Nanodevices like the Pathogen Sentinel shown below.
DNA Nanostructures and Devices: We make functional nanodevices out of DNA. We recently developed a pathogen sentinel that can detect, measure and report the presence of a variety of pathogen-related biomarkers. Billions of these sentinels can be created for pennies in a few microliters of saltwater. Even better, since they are made out of DNA they are extremely robust.
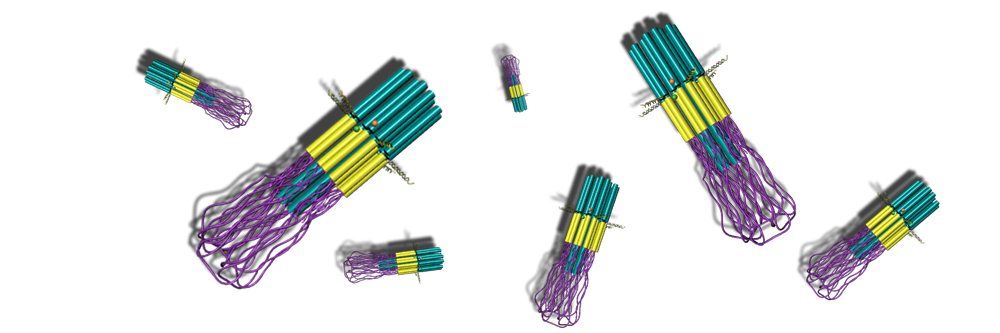
We also developed a new method for creating 2D and 3D DNA nanostructures. This method uses DNA origami as a design tool but does not require a single-stranded scaffold of biological origin. In this way, our method allows the creation of any number of DNA nanostructures with much fewer restrictions on size and, importantly, simultaneous assembly in a single reaction ("single pot" self-assembly). Creating useful machines and expanding the general method of DNA-based nanodevice construction are currently the main objectives.
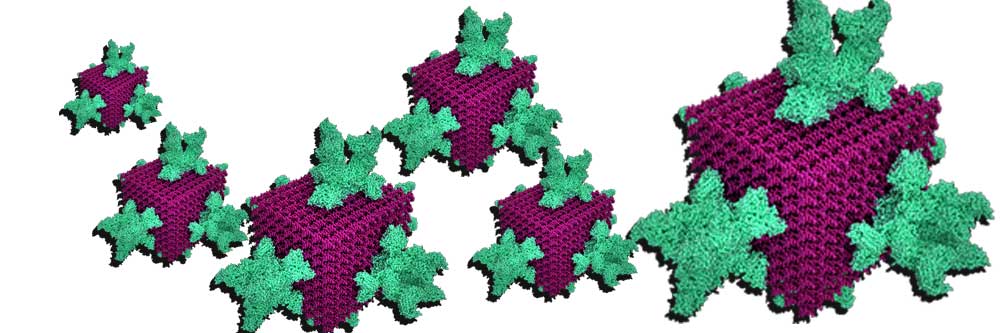
Our latest focus is on the development of a camouflaged DNA nanoparticle for targeted delivery of genes and genetic elements in living systems. As always, when it comes to nanomachinery, evolution has done every experiment for us already! Therefore, we are emulating viral strategies, synergized with DNA folding methodology, to build a custom designed DNA nanobot for genetic engineering. Welcome to the future of biomedicine : )
Recent Publications
Oh, C. Y., and Henderson, E. R. (2022) A Comparison of Methods for the Production of Kilobase-Length Single-Stranded DNA. DNA 2022, 2, 56–67. https://doi.org/10.3390/dna2010005.
Koehler, C., Mathur, D., Henderson, E. & Lutz, R. Probing the Security of DNA Origami. (2018) IEEE International Symposium on Software Reliability Engineering Workshops (ISSREW) 138–139, doi: 10.1109/ISSREW.2018.00-14.
McCloskey M. A., Mosher, C. L., Henderson, E. R. (2017) Will Artificial Trees be the Next Power Plants? Science Journal for Kids, Inc. Houston, TX, 77055, USA.
McCloskey M. A., Mosher, C. L., Henderson, E. R. (2017) Wind Energy Conversion by Plant-Inspired Designs. PLoS ONE 12(1): e0170022. doi:10.1371/journal.pone.0170022.
Mathur, Divita, and Henderson, Eric R. (2016) Programmable DNA nanosystem for Molecular Interrogation, Nature Scientific Reports, 6(27413). doi:10.1038/srep27413.
Ellis, Samuel J., Henderson, Eric R., Klinge, Titus H., Lathrop, James I., Lutz, Jack H., Lutz, Robyn R., Mathur, Divita, and Miner, Andrew S. (2014) Automated Requirements Analysis for a Molecular Watchdog Timer In Proceedings of the 29th ACM/IEEE international conference on Automated software engineering (ASE '14). ACM, New York, NY, USA, 767-778. DOI=10.1145/2642937.2643007 (Awarded the "Manfred Paul Award for Excellence in Software: Theory and Practice").
Mathur, D. and Henderson, E. (2013) Complex DNA Nanostructures from Oligonucleotide Ensembles, ACS Synthetic Biology, 2, 180-185.
R. Lutz, J. Lutz, J. Lathrop, T. Klinge, E. Henderson, D. Mathur, and D. Abo Sheasha, (2012) Engineering and verifying requirements for programmable self-assembling nanomachines, Proceedings of the Thirty-Fourth International Conference on Software Engineering (ICSE 2012, Zurich, Switzerland, June 2-9, 2012), pp. 1361-1364.
Lutz, Robyn R., Lutz, Jack H., Lathrop, James I., Klinge, Titus H., Mathur, Divita, Stull, Don M., Bergquist, Taylor G. and Henderson, Eric R. (2012) Requirements analysis for a product family of DNA nanodevices, Proceedings of the Twentieth IEEE International Requirements Engineering Conference (RE 2012, Chicago, IL, September 24-28, 2012), pp. 211-220.
People Make Science Fun!
Current Brave Souls

Eric Henderson
Oarsman | CV

Chang Oh
NanoDesigner Grad

Dhaval Ghone
NanoDesigner Undergrad

Adam Lovrien
NanoDesigner Undergrad

You?
Next NanoDesigner

You?
Next NanoDesigner

If I had my druthers I'd be inventing fun stuff, doing fun science, playing lots of fun music and writing lots of fun books - oh wait, that's what I do already! @henderbooks, erichenderson.com.
It's all about the base - and the sugar-phosphate backbone (of course). We are enamored with DNA and the magical (almost) way we can direct hundreds of little pieces to come together to make diagnostic and therapeutic nanodevices like our awesome Pathogen Sentinel (code name: OPTIMUS) and CamoNano™, a gene editor delivery nanobot!

Chang Oh

Dhaval Ghone

Adam Lovrien

Next NanoDesigner